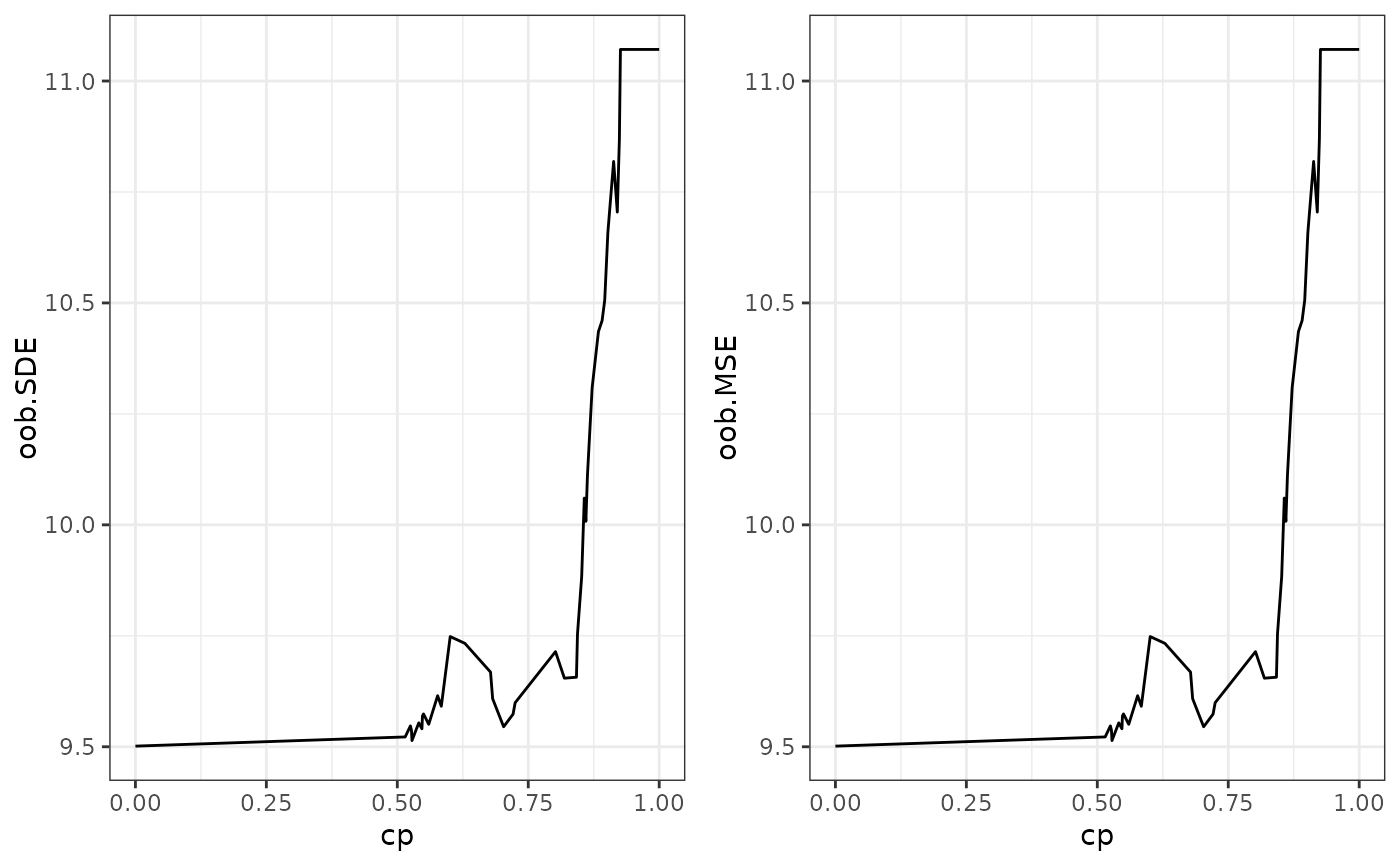
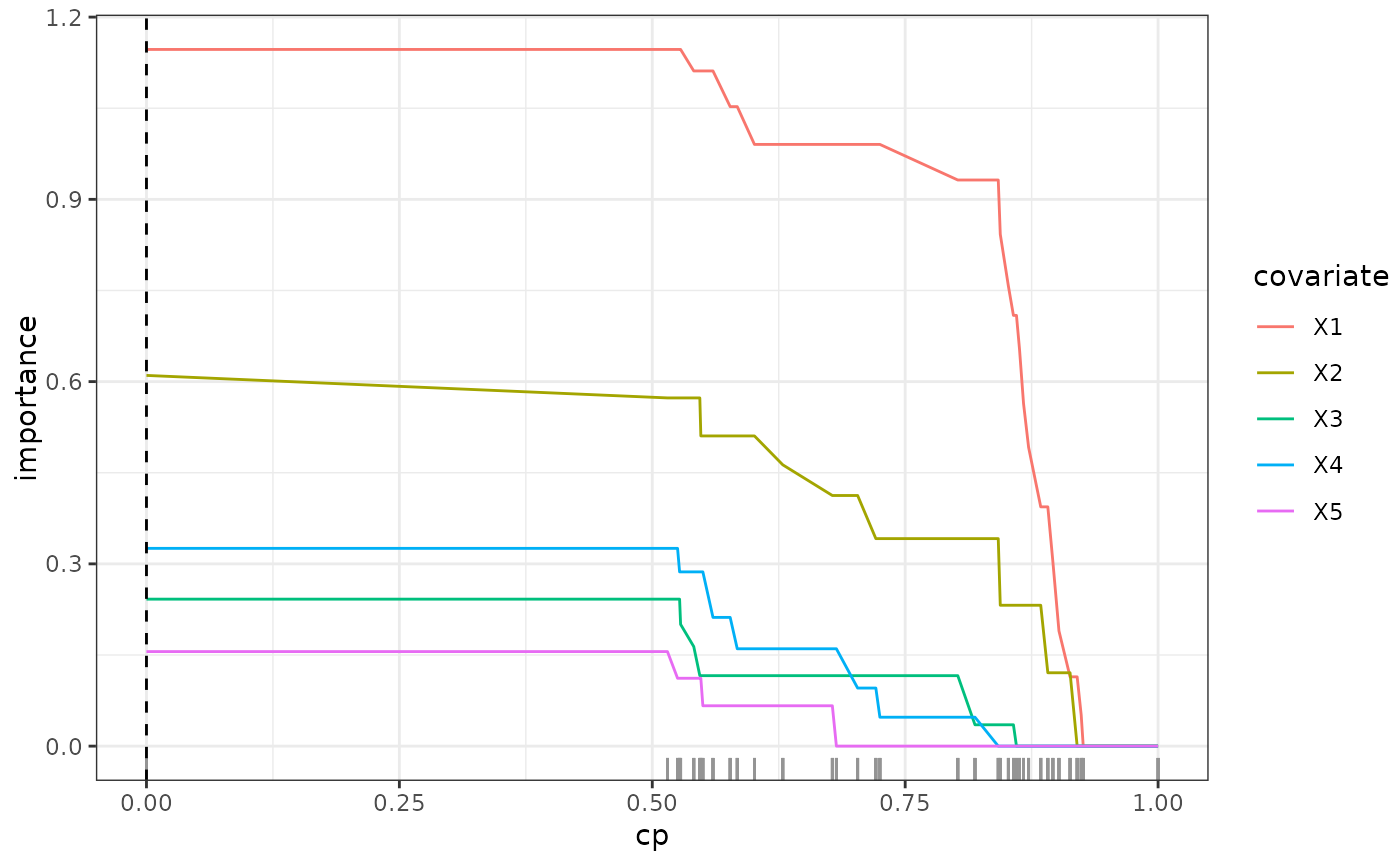
This function calculates the variable importance of an SDForest and the out-of-bag performance for different complexity parameters.
# S3 method for class 'SDForest'
regPath(
object,
cp_seq = NULL,
X = NULL,
Y = NULL,
Q = NULL,
verbose = TRUE,
mc.cores = 1,
...
)Arguments
- object
an SDForest object
- cp_seq
A sequence of complexity parameters. If NULL, the sequence is calculated automatically using only relevant values.
- X
The training data, if NULL the data from the forest object is used.
- Y
The training response variable, if NULL the data from the forest object is used.
- Q
The transformation matrix, if NULL the data from the forest object is used.
- verbose
If
TRUEprogress updates are shown using the `progressr` package. To customize the progress bar, see [`progressr` package](https://progressr.futureverse.org/articles/progressr-intro.html)- mc.cores
Number of cores to use for parallel computation `vignette("Runtime")`. The `future` package is used for parallel processing. To use custom processing plans mc.cores has to be <= 1, see [`future` package](https://future.futureverse.org/).
- ...
Further arguments passed to or from other methods.
Value
An object of class paths containing
- cp
The sequence of complexity parameters.
- varImp_path
A
matrixwith the variable importance for each complexity parameter.- loss_path
A
matrixwith the out-of-bag performance for each complexity parameter.- cp_min
The complexity parameter with the lowest out-of-bag performance.
- type
Path type