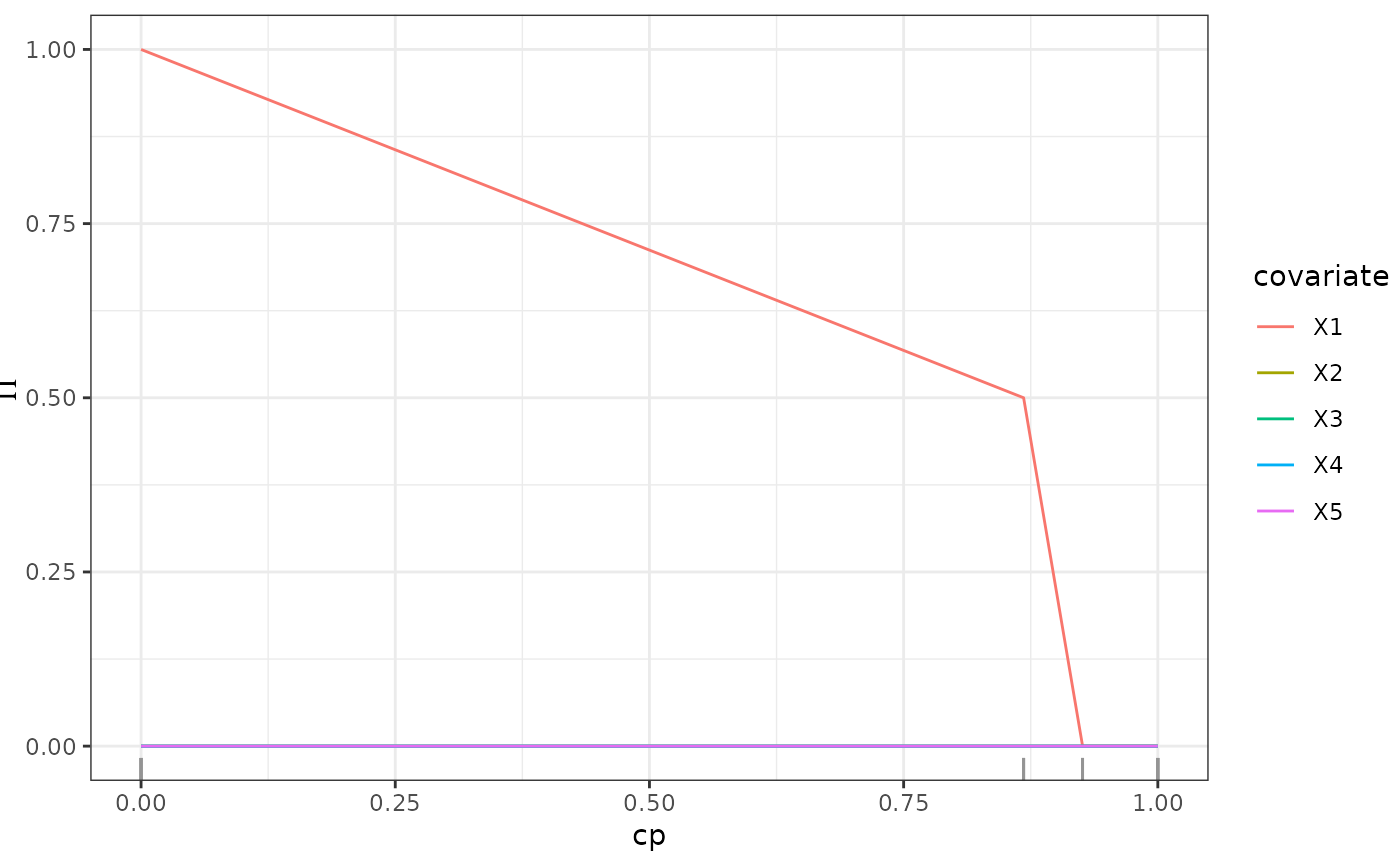
This function calculates the stability selection of an SDForest (Meinshausen and Bühlmann 2010) . Stability selection is calculated as the fraction of trees in the forest that select a variable for a split at each complexity parameter.
# S3 method for class 'SDForest'
stabilitySelection(object, cp_seq = NULL, verbose = TRUE, ...)Arguments
- object
an SDForest object
- cp_seq
A sequence of complexity parameters. If NULL, the sequence is calculated automatically using only relevant values.
- verbose
If
TRUEprogress updates are shown using the `progressr` package. To customize the progress bar, see [`progressr` package](https://progressr.futureverse.org/articles/progressr-intro.html)- ...
Further arguments passed to or from other methods.
Value
An object of class paths containing
- cp
The sequence of complexity parameters.
- varImp_path
A
matrixwith the stability selection for each complexity parameter.- type
Path type
References
Meinshausen N, Bühlmann P (2010). “Stability Selection.” Journal of the Royal Statistical Society Series B: Statistical Methodology, 72(4), 417–473. ISSN 1369-7412, doi:10.1111/j.1467-9868.2010.00740.x .